Starter Code
Get the starter code on GitHub Classroom
If you’re using Docker, you’ll want to pull the updated image: docker pull harveymudd/cs159-student:spring2020.
Overview
This lab has the following starter files:
analysis.mdngrams.pyzipf.py
You should also plan to create and submit a pdf version of your analysis, analysis.pdf.
Spacy
This week, we’ll get to use the spacy
python library for the first time. Spacy is designed to make it easy
to use pre-trained models to analyze very large sets of data. At the
beginning of the semester, we’ll be using spacy as a skeleton for
building our own NLP algorithms. Later in the semester, you’ll get a
chance to use more of the built in functionality to build larger
systems.
Zipf’s law
In this part of the lab, we will test Zipf’s law by using matplotlib, a 2D plotting library. Put your code for this part in zipf.py.
We are going to explore the relationship between the frequency of a token and the rank of that token. If you count all the tokens in carroll-alice.txt – like we did last week – we would say that the has rank of 1, and has rank of 2, to has a rank of 3, and so on, since this is the order that they appeared when we listed the tokens from most frequent to least frequent. We will use matplotlib to compare plot each token’s rank against its relative frequency.
Part 1(a)
Instead of tokenizing by hand, we will use spacy to do the tokenization. Pause now and read about space language processing pipelines.
For this part, we only want to to have spacy tokenize our text, so we will set the pipeline to []. Since some of our documents will be long, but since we’re not doing any memory-intensive processing, we will tell spacy that it’s okay to load large documents all at once. Add these lines to the top of your zipf.py file:
from spacy.lang.en import English
nlp = English(pipeline=[], max_length=5000000)
Write a function called read_one(file_path) that takes the location of a file as its input, and that returns a Counter object representing all of the lowercased tokens in the corresponding text file.
Hint: You can get the Tokens in a spacy document by iterating over the document. You can get the text of a Token (as a string) using the .text attribute of the Token object.
Hint: You may want to use the following python convention/template for opening a file and working with its contents:
with open(filename, 'r', encoding='latin1') as fp:
# do processing...
Next, write a function called read_all(dir_path, extension=None) that takes the location of a directory as its input, and returns a Counter object representing all of the text of all of the files in the corresponding directory whose extention is extension. If extension is None, then read_all should include every file in the directory. For example, read_all(/cs/cs159/data/gutenberg, ".txt"}}) should return a Counter that counts all of the tokens in all of the text files in the /cs/cs159/data/gutenberg directory.
Hints: You will want to use the os.walk function to recursively search for files in the directory. You can get a file’s extension with https://docs.python.org/3/library/os.path.html#os.path.splitext. It may also be helpful to know that two Counter objects can be added together to create a new Counter object!
Part 1(b)
Let f be the relative frequency of a word (e.g. if the occurs 1642 times out
of 35652 tokens, then its relative frequency is 1642/35652 =
0.04606), and let r be the rank of that word.
To visualize the relationship between rank and frequency, we will
create a log-log
plot of r (on
the x-axis) versus f (on the y-axis). We will use the pylab
library, part of matplotlib. By default, matplotlib will try to open a window to display figures as soon as they’re created. That won’t work over ssh (unless you’re using window forwarding) or in Docker, but we can stop matlab from opening windows by changing which backend it uses. This needs to be done before we import pylab:
import matplotlib
matplotlib.use('Agg')
from matplotlib import pylab
You will write a function called do_zipf_plot that takes two parameters:
- A
Counterobject with the counts of words from one or more files. - A
stringlabel that can be used to title the figure.
The starter code includes two implemented functions that call do_zipf_plot:
plot_one, which callsread_oneanddo_zipf_plotto generate a visualization of data from one file, andplot_all, which callsread_allanddo_zipf_plotto generate a visualization of data from an entire directory.
Your do_zipf_plot function should start by creating a figure object:
fig = pylab.figure()
It should then use the Counter argument to create data in the right form for a call to pylab.loglog
Be sure to label your plot with xlabel, ylabel, and suptitle. Add a legend to the lower left of the plot, and then save the resulting figure:
pylab.savefig('zipf_{}.png'.format(label))
pylab.close()
Before moving on, confirm that calling plot_one('carroll-alice.txt') generates a plot that matches:
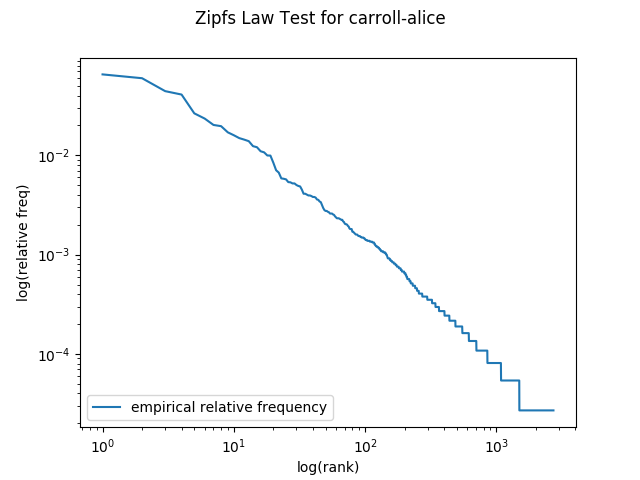
Part 1(c)
Now we can test how well Zipf’s law works. Read Wikipedia’s article on Zipf’s law. In summary, Zipf’s law states that \(f \propto \frac{1}{r}\), or equivalently that \(f = \frac{k}{r}\) for some constant factor \(k\), where \(f\) is the frequency of the word and \(r\) is the rank of the word. Following Zipf’s law, the 50th most common word should occur with three times the frequency of the 150th most common word.
Add to your do_zipf_plot function so that in addition to plotting the empirical rank vs frequency data, it also plots the expected values using Zipf’s law. For the constant \(k\) in the formulation of Zipf’s law above, you should use \(\frac{T}{H(n)}\) where \(T\) is the number of word tokens in the corpus, \(n\) is the number of word types in the corpus, and \(H(n)\) is the \(n^{th}\) harmonic number. The number of tokens is the total number of words in the document. The number of types is the total number of unique words in the corpus.
Use this function to compute harmonic numbers. It’s included in your starter code:
def H_approx(n):
"""
Returns an approximate value of n-th harmonic number.
http://en.wikipedia.org/wiki/Harmonic_number
"""
# Euler-Mascheroni constant
gamma = 0.57721566490153286060651209008240243104215933593992
return gamma + math.log(n) + 0.5/n - 1./(12*n**2) + 1./(120*n**4)
To plot a second curve, you will add a second pylab.loglog(...) line after you plot the empirical frequency data. Be sure to label each data line so that your lengend will be informative!
Questions
Answer the following questions in the corresponding parts of analysis.md. Be sure to include plots you get for each part.
-
Zipf’s Law and Alice: How well does Zipf’s law approximate the empirical data in
carroll-alice.txt? -
Zipf’s Law and Other Texts: Repeat the previous question for a few other texts included in the
/cs/cs159/data/gutenbergdirectory. Are the results consistent? - Zipf’s Law and All Texts: Repeat the Zipf’s law experiment with all of the text from all of the files in
/cs/cs159/data/gutenberg.- How many tokens are in this new corpus?
- How does this plot compare with the plots from the smaller corpora?
-
Zipf’s Law Discussion: Does Zipf’s law hold for each of the plots your made? What intuitions have you formed? Does the length of a document have an impact on how well it does or does not follow Zipf’s Law? What else do you notice?
- Zipf’s Law for Random Text: Generate random text, e.g., using
random.choice("abcdefg... "), taking care to include the space character. You will need toimport randomfirst. Use the string concatenation operator to accumulate characters into a (very) long string. Then tokenize this string, and generate the Zipf plot as before, and compare the plot to the ones you got from “real” English data. What do you make of Zipf’s Law in the light of this? (Source: Exercise 23b, Bird, Klein and Loper, 2009)
NGrams
This week, we learned about the problem of 0’s in language modeling. If you were to train a unigram language model on the fiction category of the Brown corpus and then try to calculate the probability of generating the editorial category, you’d end with a 0 probability because there are words that occur in the editorial category that don’t appear in the fiction category (e.g. “badge”).
In this part of the assignment, we’ll explore that problem in more depth. We’ll also start working with the dataset that you’ll use for your final project.
You should put your code for this part in ngrams.py. Add the same lines to the top of this file for working with spacy that you have in your zipf.py.
Part 2a: Extracting data from XML files
A sample of the dataset we’ll use for your final project
is in /cs/cs159/data/semeval. There’s a single XML file that contains
all of the articles we’ll look at this week. Each article has a hyperpartisan attribute that indicates whether is it an example of hyperpartisan news: “true” or “false”.
Eventually, you may want to use a much larger (~3.6G) data file related to this one in your final project. Loading an xml file that big into memory is a recipe for trouble, though, so it’s best not to store the whole
thing in memory at once if we can help it. Fortunately, the lxml library gives us a way to
iteratively parse through an xml file, dealing with one node at a
time. Here’s sample code that opens a file called myfile.xml and
call a function called my_func on every article node:
from lxml import etree
fp = open("myfile.xml", "br")
for event, element in etree.iterparse(fp, tag=("article",)):
my_func(element)
element.clear()
The starter code has a generator function called do_xml_parse() that uses lxml.etree to yield one node at a time. Look at that code and make sure you can explain how each line of it works before you move on.
Write a function called get_unigrams that takes as input a spacy Document, and returns a list of all of the unigrams in the document. get_unigrams should also take an optional argument do_lower whose default value is True. That argument should determine whether the text of each token is lowercased before returning the unigrams. For all of the analysis in this lab, you should lowercase the tokens unless told otherwise.
Next, write a function called get_articles(args, attribute, value) that returns a Counter. get_articles will use do_xml_parse to iterate through all of the articles in args.articles. For each article whose attribute attribute has the value value, it will call get_unigrams on the text of the article. For example, get_articles(args, 'hyperpartisan', 'true') will call get_unigrams once for every article whose hyperpartisan attribute is true.
Hint If you have an article element called article, you can access all of the text children of that element with article.itertext().
Hint: Some of the articles contain html entities. You can unescape those by importing the html module and calling html.unescape on the articles’ text before creating your spacy docs.
Stop now and confirm that if you call get_articles on the semeval-sample.xml file for the case where hyperpartisan=true, you get the following counts:
the: 10631
california: 34
zero: 11
We are interested in knowing how many of the unigrams in one category of text (e.g., hyperpartisan='true') are not in another category of text (e.g., hyperpartisan='false'). Over the course of this assignment, you will explore several ways of grouping the text, so we’ll want to carefully organize our code for reusability. In the rest of this writeup, we’ll refer to the set of data we generate counts from as the training set, and the set of data that we check for zeros using those counts as the test set.
Write a function called compare(train_counter, test_counter, unique=False). The three arguments to compare should be:
train_counter: ACounterobject representing counts from the training settest_counter: ACounterobject representing counts from the test setunique: A boolean indicating whether to count zeros for tokens (unique=False) or types (unique=True)
compare should return two numers:
- The count of (tokens/types) in the test set that have a zero count in the training set
- The total number of (tokens/types) in the test set
Confirm that if you call compare(Counter([1,2,3]), Counter([3,4,4]), unique=True) you get (1,2), and if you call compare(Counter([1,2,3]), Counter([3,4,4]), unique=False) you get (2,3).
The given code has a function called do_experiment that calls get_articles twice (once for the training data, once for the test data), and then prints the results from compare as a markdown table. Read through that function now and make sure that you understand it, since you will add it it later in the lab.
Questions
- What percentage of the tokens that appear in the ‘hyperartisan (True)’ articles don’t appear in the ‘neutral (False)’ articles?
- What percentage of tokens that appear in the ‘neutral (False)’ articles don’t appear in the ‘hyperpartisan (True)’ articles?
- What if you look at types instead of tokens?
- Are you surprised by these results? Why or why not?
Part 2b: Bigram analysis
What happens when you move to higher order n-gram models like bigrams and trigrams?
Write a function called get_bigrams that takes as input a spacy Document, and returns a list of all of the bigrams in the document.
Write a function called get_trigrams that takes as input a spacy Document, and returns a list of all of the trigrams in the document.
Hint: Don’t try to manually generate the bigrams and trigrams. Instead, use your get_unigrams function along with python’s built-in zip function!
Modify your get_articles function so that it also creates and returns a Counter of bigrams and a Counter of trigrams. Then modify do_experiment so that it also calculates and reports statistics about bigram and trigram zeros.
Questions
-
What percentage of the bigrams (tokens, not types) that appear in the ‘hyperpartisan’ articles don’t appear in the ‘neutral’ articles? What percentage of the bigrams that appear in the ‘neutral’ articles don’t appear in the ‘hyperpartisan’ articles? Are you surprised by these results? Why or why not?
-
What percentage of the trigrams (tokens, not types) that appear in the ‘hyperpartisan’ articles don’t appear in the ‘neutral’ articles? What percentage of the trigrams that appear in the ‘neutral’ articles don’t appear in the ‘hyperpartisan’ articles? Are you surprised by these results? Why or why not?
Part 2c: Train on equal-sized “chunks”
Instead of training on one category of articles and testing on another, suppose we break each of the categories in half. Then we could train on half of the hyperpartisan articles and half of the neutral articles, and test on the other half.
In the sample data you used above, each of the articles has an attribute randomchunk that assigns it to either chunk a or chunk b.
You shouldn’t need to write much (any!) code here. Instead of calling do_experiment on the hyperpartisan attribute, you can now call it on the randomchunk attribute.
Questions
- Try training on randomchunk
aand testing on randomchunkb.. Then train on randomchunkband test on randomchunka. How are your results different from the previous question? Why? Your writeup should include a table of your results, which you can generate with your expandeddo_experimentfunction from above. Report percentages, not raw counts.
Evaluating your system in this way is called cross-validation. In this case, since you are breaking your data into 2 distinct test sets, you are performing 2-fold cross-validation.