
To use Jane, begin by downloading the .zip file available from the main page. Once you have unzipped it, there will be a jar file in the Jane folder. To run Jane, use the command java -jar Jane.jar. On some machines, you can simply click or double-click on the Jane.jar icon. (If Jane crashes when you run on a large problem, it is almost certainly because Java ran out of memory. Use the command line option -Xmx512m in this case.)
There is also a command-line version of jane included in the jar which can be invoked with java -cp Jane.jar Jane treefile. This version of Jane is particularly useful for conducting large-scale experiments controlled by scripts. More details on this option are given at the end of this tutorial.
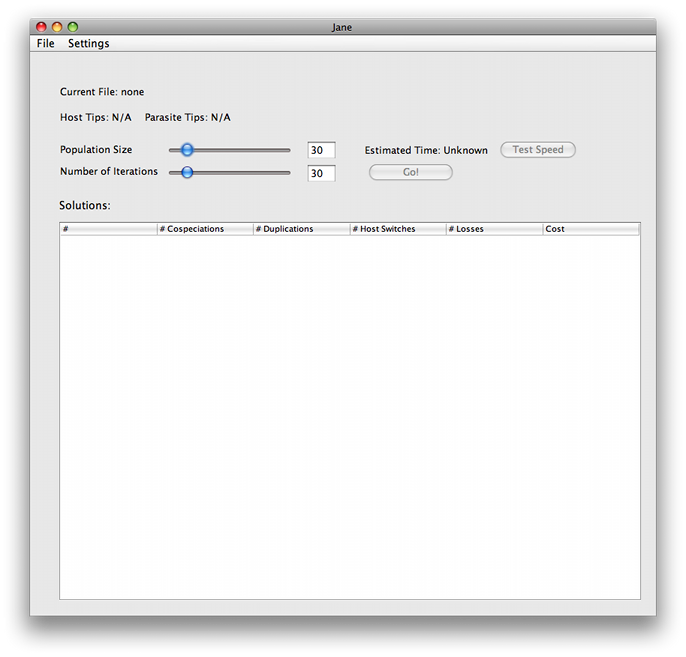
Next, open a tree file by going to File > Open Tree. Jane supports two formats: .nex files, formatted in the style used by TreeMap, and .tree files, formatted in the style used by Tarzan. A more exact specification of file formats used by Jane, as well as some example files can be found here.
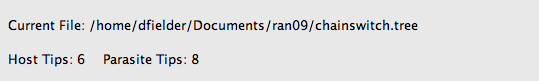
Once you open a tree file, its filename and a count of host and parasite tips will appear at the top of the window.

Jane has several configuration options that determine how well it reconciles the trees, and what criteria to use.
 (command line option -p size)
(command line option -p size) (command line option -i size)
(command line option -i size)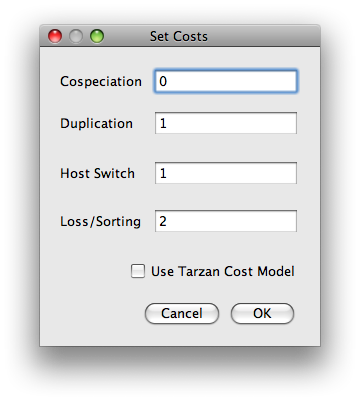
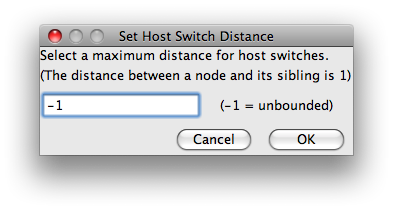
Once you are done configuring Jane, you should click the Go button. Jane will then run through the genetic algorithm. Those solutions will appear in the list at the bottom of the screen
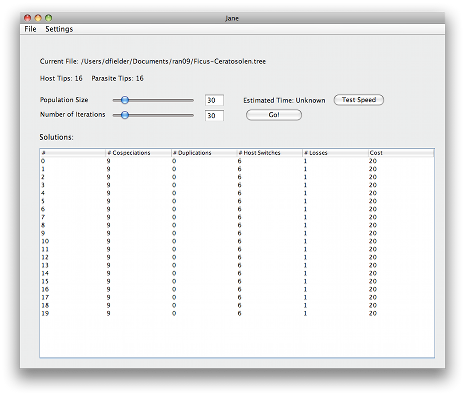
Each of these solutions is a relative timing of the vertices in the
host tree. It is likely that many of them will contain similar optimal
solutions.
Double-click on one of the solutions to bring up a solution viewer.
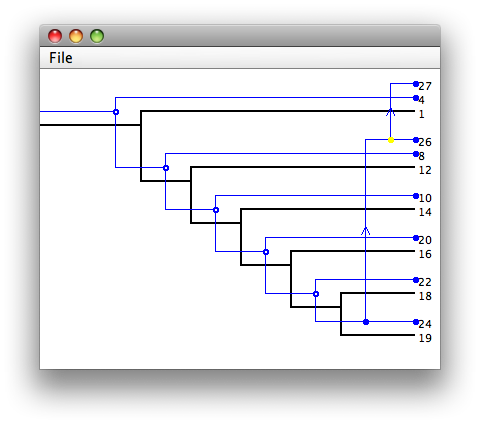
Within the solution viewer, there are four types of events
| Cospeciation | 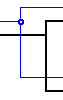 |
| Duplication | 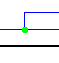 |
| Host Switch | 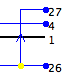 |
| Loss | 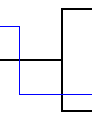 |
Note: On Windows and Linux machines, the small "hole" in the circle for a copseciation node is very small and may be hard to see on some displays. However, note that a cospeciation always has two vertical segments emanating from it, which clearly distinguishes it from a duplication which always has one vertical and one horizontal emanating segment.
Notice that each type of event corresponding to a vertex in the
parasite tree is marked with a colored circle. These circles
indicate the existence of other possible locations for the
association. A green circle means there is a location of lower cost
where the parasite vertex could be mapped. A yellow circle means that
there is a location of equal cost, and red circle means that any other locations that may exist are of higher cost. When dragging an association, segments of the host tree will light up colors corresponding to the cost of moving the association to that edge or vertex.
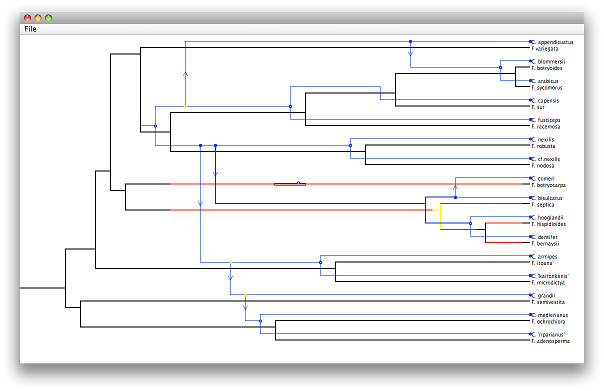
If a timing is of interest, you can save it (but not any changes made to the parasite tree within that timing) by going to File > Save Timing inside the solution viewer window. Then, any time that tree is loaded in the main window, you can go to File > Load Timing, open the timing, and it will be added back into the list of solutions at the bottom of the window. Command-line users can save the best timing found by using the -o switch with the name of a file to store the timing in.
It's worth noting that the timing files are written in a human-readable format, so they can be modified if you want to explore a specific timing.
Jane can be invoked from the command-line rather than by launching the
graphical user interface as follows:
java -cp Jane.jar Jane treefile
where treefile is the file name to be used. A full listing
of the command-line options can be found as follows:
jave -cp Jane.jar Jane -help
Finally, for very large tree files and timing files,
Jane's interactive viewer may be
too slow or may exceed the memory of the computer. Therefore, a
version of Jane with a more efficient
non-interactive viewer is available by invoking:
java -cp Jane.jar SimpleViewer
This version has all of the command-line options of Jane. In
addition, there is also a -o switch that outputs an image (in png
format) of the solution.
Back to Jane homepage