Biological Plausibility of Neural
Network Models
Irina Rabkina and Jessica Schroeder
Overview
Throughout the course of our neural networks class we learned about numerous different neural network models, all of which are useful in computer science applications such as classification problems and unsupervised learning. However, as neural networks are often used to model the biological system off which they are based, we aimed to examine the actual biological plausibility of several of the models we discussed in class. We researched a few controversial examples of such models, and present the arguments for and against their biological relevance and discuss the importance of these considerations on this page.
Main Questions
We had a few questions we wanted to keep in mind throughout our investigation:
How biologically relevant are current models?
We wanted to determine the extent to which ANY current models are really biologically relevant.
Are some models more biologically relevant than
others?
We wanted to examine the difference in biological plausibility between different models.
How biologically relevant do models need to be in
order to be trusted in scientific studies of the brain?
Humans consider experimenting on human brains immoral, so instead researchers use monkeys, rodents, and even insects as models. Clearly, there’s a point at which we say “we know this isn’t a perfect model of the human brain, but it’s close enough to be helpful.” Where is this point for artificial neural networks? This is a question scientists will be debating for decades—we’re not going to be able to answer it ourselves—but we thought it important to consider.
Is biological relevance of the model more important
than the model’s performance?
If a researcher is faced with the choice between a biologically relevant network that does not produce consistent results and a less biologically relevant model that exhibits the desired behavior, does she need to use the biologically relevant model for biological studies? Or is it legitimate to say that if it’s performing well, it can give us insights into the behavior it’s modeling?
Neural Network Models: General
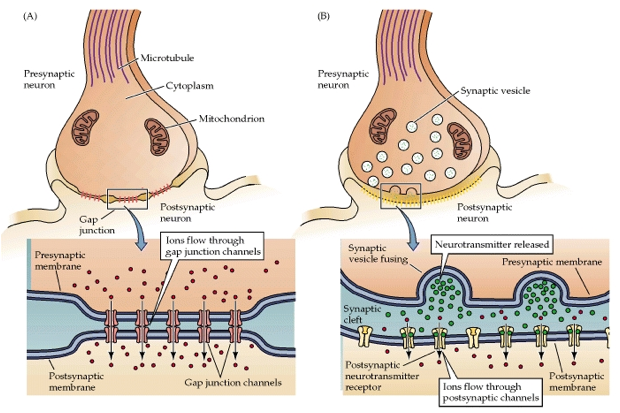
A diagram of a chemical synapse, the most prevalent type in the brain, which models fail to fully mimic (Clark, 2013).
Although connectionist neural network models often
provide very good fits to experimental data, Sharkey and Sharkey (1998) argue
that many aspects of such models are biologically problematic. For one,
the programmer chooses aspects such as the basic architecture, the learning
technique, the learning rate, and even the data representations. Because
of this, models generally are tweaked to fit the data, instead of staying true
to any aspects of human learning. Another problem is the fact that in
computational models, neurons are all the same, while the brain has many different
types of neurons—each with a different combination of the many different
neurotransmitters that exist in the brain—that work together to cause
behavior and cognition. Computational models also only model electrical
synapses, which are rare; the brain is primarily made of chemical synapses,
which can have longer-lasting effects than electrical synapses (Spencer, 2009).
In addition, biological neural networks have huge numbers of neurons,
while computational ones have orders of magnitude fewer, and biological
networks are able to perform much more quickly than artificial neural networks.
Finally, the difference between neurons and computer chips in terms of
proneness to malfunction is approximately 1 to more than 109. This makes sense—we don’t want our
personal computers malfunctioning—but the fact that our models fail to
exhibit the randomness, and therefore the necessary robustness, that the brain
must exhibit is a huge difference. We
therefore lose nuances of the brain when trying to use artificial neural
network models to approximate it.
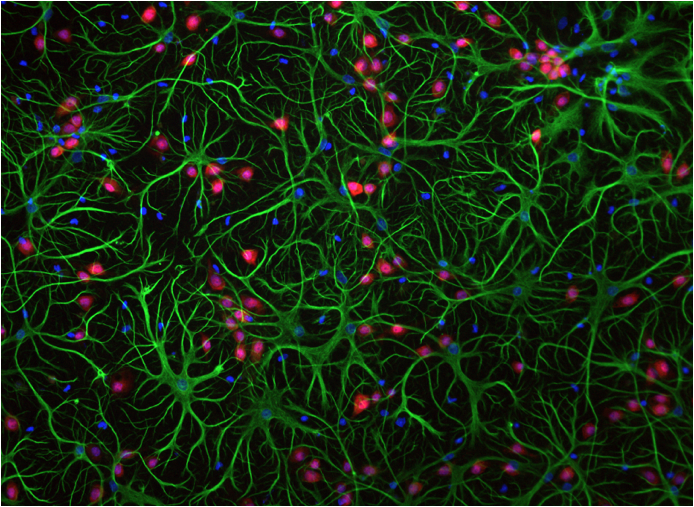
Glial cells (green and red) and neurons (blue). Schlesinger, 2011.
One last problem we want to point out with the
concept of modeling the human brain is the degree to which we simply do not
have the knowledge we need to accurately model it. Even biological phenomena that
scientists consider fairly well-characterized are not completely understood. The best example of this lack of
knowledge is the fact that we only ever model neurons, not glial cells. Glial cells make up about 90% of the
cells in the human brain. In the
image above, the green and red cells are glial cells; only the blue cells are
neurons. We don’t talk much about
glial cells simply because we’re still not quite sure exactly what they
do. Their name literally means
“glue,” and came about because we thought their only function was structural
support. However, now know that they’re
involved in the breakdown of certain neurotransmitters, and allow for
transmission of action potentials over longer axons. The fact that we’re ignoring the
existence of these cells in our models shows the extent to which we are NOT
modeling the brain.
Hopfield Networks
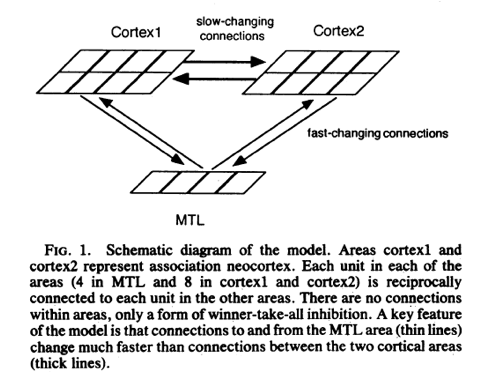
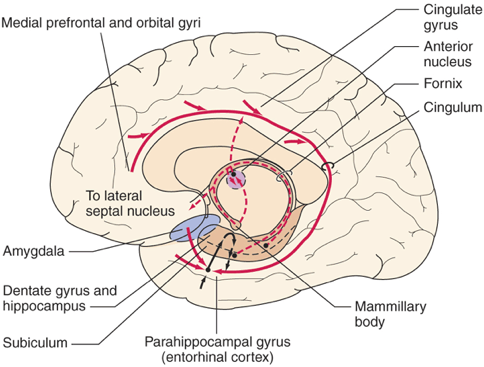
A Hebbian-Based Network Model of the Medial Temporal Lobe Image reference: Thebrainlabs.com
(Alvarez & Squire, 1994).
Hopfield models are used fairly
frequently in connectionist models of cognitive function. For example, such
networks have been used in models of memory disorder in diseases such as
Alzheimer’s disease (Zhao et al., 2010) and memory consolidation in the medial
temporal lobe (Alvarez & Squire, 1994). Hopfield networks rely on Hebbian
learning rules to create the connections between nodes of the network. Hebbian learning rules are based off of biological phenomena
such as associative learning: in associative learning, neurons that fire
together have strengthened connectivity; the same holds true in Hebbian learning, as the connection between the
postsynaptic and presynaptic neurons strengthens if they fire together.
However, many aspects of Hebbian learning
rules, such as the requirement for symmetry, are biologically unfeasible (Mazzoni et al., 1991). In addition, Hopfield networks
simply fall into a stored pattern given an input; they never update. As biological
neural networks, especially those involved in memory processes, are
continuously updating, a static model is not particularly biologically
relevant.
Bolzmann Machines and Deep Learning
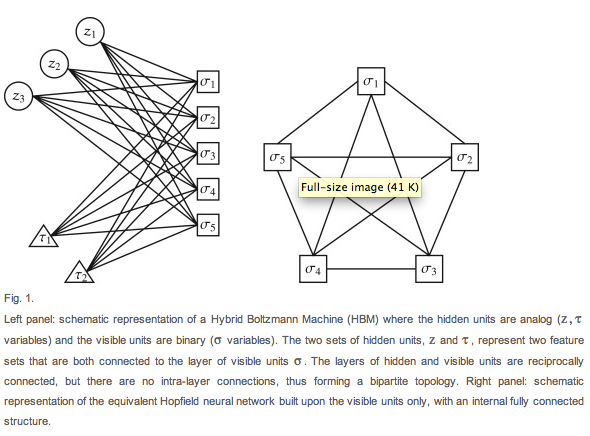
Boltzmann Machine (left) vs Hopfield Networks (right) (Barra et al., 2012).
Boltzmann Machines can be used to model many of
the same probability distributions as Hopfield Networks, with a lower space
requirement but slightly higher time requirement (Barra, Bernacchia,
Santucci & Contucci
2012). From a neural modeling perspective, this is important as it gives a
biologically more plausible alternative to Hopfield Networks.
Furthermore, O'Reilly
(1998) claimed that “The form
of synaptic modification necessary to implement [Boltzmann Machines] is
consistent with (though not directly validated by) known properties of
biological synaptic modification mechanisms,” as Boltzmann Machines update
stochastically. In addition, in proposing a parallel search
algorithm adapted from statistical mechanics, Ackley, Hinton, and Sejnowski suggested that the algorithm is biologically
plausible as a model for classification in the brain because computing units
are small and connections hold numeric values (1985). Restricted Boltzmann
Machines, which build on the Boltzmann Machine, are now commonly used in deep
learning--a process that has been likened to various types of learning in the
biological brain (Utgoff and Stracuzzi,
2002). All this suggests that Boltzmann Machines are more biologically plausible
than Hopfield Networks. However, Boltzmann Machines do also have symmetric
weights, so they are far from perfect.
Self-Organizing Maps
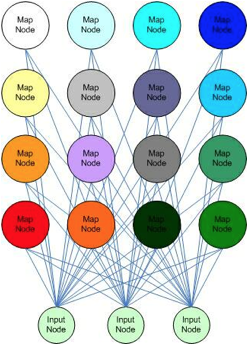
Self-Organizing
Map based on Color
(Chestnut, 2004).
Self-Organizing Maps are by far the most
biologically relevant models we examined in our investigation. Self-organizing maps have been proposed
as bases for a number of neurological phenomena. They have been used to
model everything from lower-level mechanisms such as the organization of the
striate cortex (von der Malsburg, 1973) to higher-level concepts such
as pathology found in autism (Noriega, 2007). However, some dispute the
biological plausibility on concepts such as Euclidian distance and the global
supervision necessary to select neighborhoods (Miikkulainen,
1991). In addition, many
self-organizing maps have no concept of time. They simply take all the
inputs and create the maps in accordance with space. Chappell and Taylor
(1993) argue that for a self-organizing map to be at all biologically relevant
it must be temporally sensitive as well, because human learning depends so much
on time. It is important to note, however, that much of human memory is
‘self-organizing;’ for example, grid cells within the entorhinal
cortex organize into hexagonal patterns as one learns about spatial navigation
(Mhatre et al., 2012), and language acquisition can
be modeled using self-organized neural networks (Li et al., 2004). They
therefore seem to have some biological relevance, especially when augmented
with a temporal sensitivity.
In addition, the basic features of Self-Organizing
Maps are more plausible than many other models. For example, they have a high degree of
connectivity, and they follow Hebbian learning rules—neurons
that fire together get stronger synaptic strength between them (Spitzer, 1995). This is a phenomenon observed in human
neuroscience and is an important aspect in any model of the brain. In addition, two inputs that are similar
will be closer together than two inputs that are very different. As the human homunculus is organized in
a similar matter—with the area of the brain that encodes signal for the
fingers on the left hand being located near the area of the brain that encodes signal
for the left palm, for example—this seems to be very biologically
consistent. Lastly and most
importantly, Self-Organizing Maps use unlabeled data, whereas many neural
network models rely on labeled data.
The human brain does sometimes get feedback when it makes a mistake, but
the feedback is often much more vague and delayed than feedback in models that
use labeled data. The brain rarely
gets any feedback so clear-cut as “I outputted 00101 and should have outputted
00110, so I should now account for that error,” and models that don’t rely on
such feedback are more relevant than models that do.
Ongoing Projects
Several
major projects are looking to model the human brain more closely and/or
understand it well enough to create such models. A few of these are described
below.
NeuroGrid
NeuroGrid is a neuromorphic
circuit built by Benjamin et al. (2014) that seeks to model processes of entire
brain. It can perform many of the same computations as the brain, but it is
much less efficient in terms of time, memory, and power. It is also much larger
than the human brain. The scientists are seeking to update the circuit with
newer technology in order to improve its performance; however, new technology
may not be enough—we simply do not understand what makes our brains as
efficient as they are, so recreating this efficiency at this point in time is
nearly impossible.
The Virtual Brain
The Virtual
Brain is an open-source project that allows users to model structural and
functional connectivity. The connectivity is based off of DTI and fMRI data
from real human participants. This is, however, very much a work in progress
and is not yet particularly accurate (Jirsa et al.,
2010).
The Human Connectome Project
The Human Connectome Project is a long-term collaboration between
several universities in the United States and abroad that seeks to accurately
model the structure and function of the human brain. They have collected structural
data using HARDI, and R-fMRI and functional data using fMRI, EEG, MEG for
models of function from hundreds of participants (Toga et al., 2012). This data
will be combined to learn more about the brain; hopefully this will allow
scientists to create a full working model in the near future.
References
·
Ackley, D. H., Hinton, G.
E. and Sejnowski, T. J. (1985), A Learning Algorithm
for Boltzmann Machines. Cognitive Science, 9: 147–169.Alvarez, P. &
Squire, L. (1994). Memory consolidation and the medial temporal lobe: A simple
network model. PNAS. 91. p.7041-7045.
·
Barra, A., Bernacchia, A., Santucci,E. &
Contucci, P. (2012).On the equivalence of Hopfield
networks and Boltzmann Machines.Neural Networks, 34.
·
Benjamin, B.V.; Gao, P.; McQuinn, E.; Choudhary, S.; Chandrasekaran,
A.R.; Bussat, J.-M.; Alvarez-Icaza,
R.; Arthur, J.V.; Merolla, P.A.; Boahen,
K., (2014) "Neurogrid: A Mixed-Analog-Digital
Multichip System for Large-Scale Neural Simulations," Proceedings of the
IEEE , vol.PP, no.99, pp.1,18
·
Chappell, G. J., &
Taylor, J. G. (1993). The temporal Kohønen map.
Neural networks, 6(3), 441-445.
·
Chesnut, Casey. "Self Organizing Map AI for Pictures." (2004).
·
Clark, Sean. “KIN-450 Neurophysiology Wiki.” 2013. Accessed 20 April 2014.
·
Dyer, Michael G. "The
promise and problems of connectionism." Behavioral and Brain Sciences
11.01 (1988): 32-33.
·
Hoffman RE (1997) Neural
Network Simulations, Cortical Connectivity, and Schizophrenic Psychosis MD
Computing 14:200-208
·
Jirsa, V. K., Sporns, O., Breakspear,
M., Deco, G., & McIntosh, A. R. (2010). Towards the virtual brain: network
modeling of the intact and the damaged brain. Arch. Ital. Biol,
148(3), 189-205.
·
Li, P., Farkas,
I., & MacWhinney, B. (2004). Early lexical
development in a self-organizing neural network. Neural Networks, 17(8),
1345-1362.
·
Mazzoni, P., Andersen, R. A. & Jordan, M. I. (1991). A more biologically
plausible learning rule for neural networks. Proceedings of the National
Academy of Sciences, 88(10), 4433-4437.
·
Mhatre, H., Gorchetchnikov, A., & Grossberg, S. (2012). Grid cell hexagonal patterns formed
by fast self‐organized learning within entorhinal cortex.
Hippocampus, 22(2), 320-334.
·
Miikkulainen, Risto. "Self-organizing process based
on lateral inhibition and synaptic resource redistribution." Proceedings
of the International Conference on Artificial Neural Networks. 1991.
·
Noriega, Gerardo.
"Self-organizing maps as a model of brain mechanisms potentially linked to
autism." Neural Systems and Rehabilitation Engineering, IEEE Transactions
on 15.2 (2007): 217-226.
·
O'Reilly, R. C. (1998). Six
principles for biologically based computational models of cortical cognition.
Trends in cognitive sciences, 2(11), 455-462.
·
Sharkey, A., & Sharkey,
N. (1998). Cognitive modeling: Psychology and connectionism. In M. Arbib (Ed.),The Handbook of Brain Theory and Neural
Networks (1 ed., pp. 200-203). Cambridge, Massachusetts: MIT Press.
·
Schlesinger, Mark
(2011). The role of Glial Cells and
Neuroplasticity in Pain Reception.
Schlesinger Pain Center.
·
Spencer KM (2009) The
Functional Consequences of Cortical Circuit Abnormalities on Gamma Oscillations
in Schizophrenia: Insights from Computational Modeling. Front Hum Neurosci.
3:33.
·
Spitzer, M. (1995). A neurocomputational approach to delusions.Comprehensive
psychiatry, 36(2), 83-105.
·
Toga, A. W., Clark, K. A.,
Thompson, P. M., Shattuck, D. W., & Van Horn, J. D. (2012). Mapping the
human connectome. Neurosurgery, 71(1), 1.
·
Utgoff, P.E. and Stracuzzi, D.J. “Many-layered
learning.” Neural Computation 14.10 (2002).
·
von der Malsburg,
Chr. "Self-organization of orientation sensitive cells in the striate
cortex." Kybernetik 14.2 (1973): 85-100.
·
Zhao, Wangxiong,
Qingli Qiao, and Dan Wang.
"A Hopfield-like hippocampal CA3 neural network model for studying
associative memory in Alzheimer's disease." Neural Regeneration Research
5.22 (2010).